Hello, is it possible to measure the volume of a bone defect in a CT dataset with the ImFusion Suite software? And if so, how do I do that?
Do you have the ImFusion Labels software?
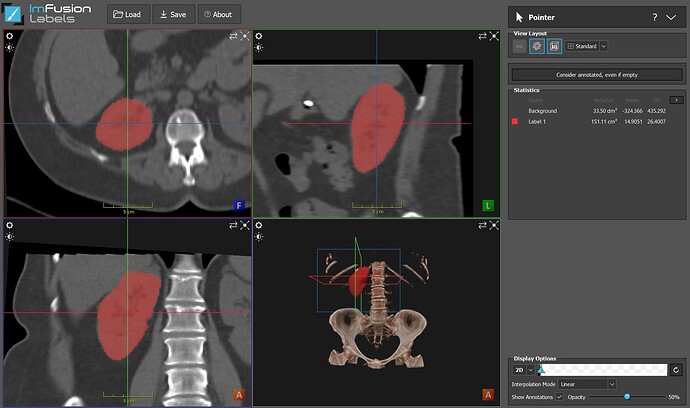
Measuring volumes of structures is something that can be easily done in this software: just annotate it (using our 3D Brush for instance) and the Statistics will be shown in the Pointer sub-menu.
Hi,
alternatively in the ImFusion Suite you can reach a similar functionality using the Segmentation > Labeling algorithm: It enables you to perform a similar brush-based labeling of the bone and offers the Statistics button which will print out the label statistics in the log window on the bottom, for instance:
Calculating statistics on the CPU:
2 labels comprising 284546 of 16777216 voxels (1.69603%)
1: 191194 voxels (1.13961%), 5493.74 mm3, mean -95.8895, std 94.3366, min -660, max 525, snr -1.01646
2: 93352 voxels (0.556421%), 2682.36 mm3, mean 122.023, std 688.314, min -1024, max 2859, snr 0.177278
If you already have an existing label map, you could also run the Segmentation > Structures View algorithm and then right click on the label and click “Show Statistics” opening a new window with that information.
Cheers,
Christian
Thank you for the quick responses. Unfortunately I don’t have the ImFusion Labels software. In Imfusion Suite, the only segmentation algorithms available to me are Spline Slicing, Superpixels Segmentation and Vesselness. What is the reason for this?
Hi,
this sounds as if you are lacking the Segmentation Plugin/Module.
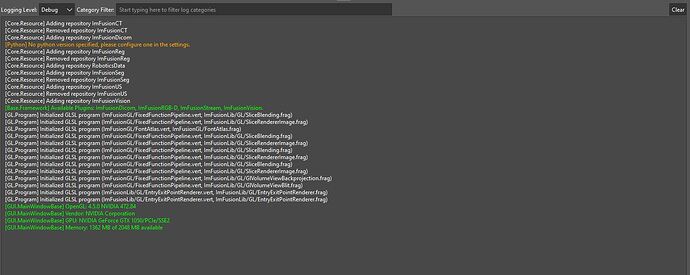
Can you please check what the log window prints on application start? There should be a line where it lists the loaded plugins like [Framework] Available Plugins: ImFusionDicom, ImFusionML, ImFusionReg, ImFusionSeg, TorchPlugin.. The Labeling/Measurement facilities are part of the ImFusionSeg plugin.
Cheers,
Christian
No, ImFusionSeg does not apppear in the list of available plugins. Since I see Vision/RGB-D in there instead I assume that your vision license does not include the segmentation plugin and it is therefore disabled.
Cheers,
Christian